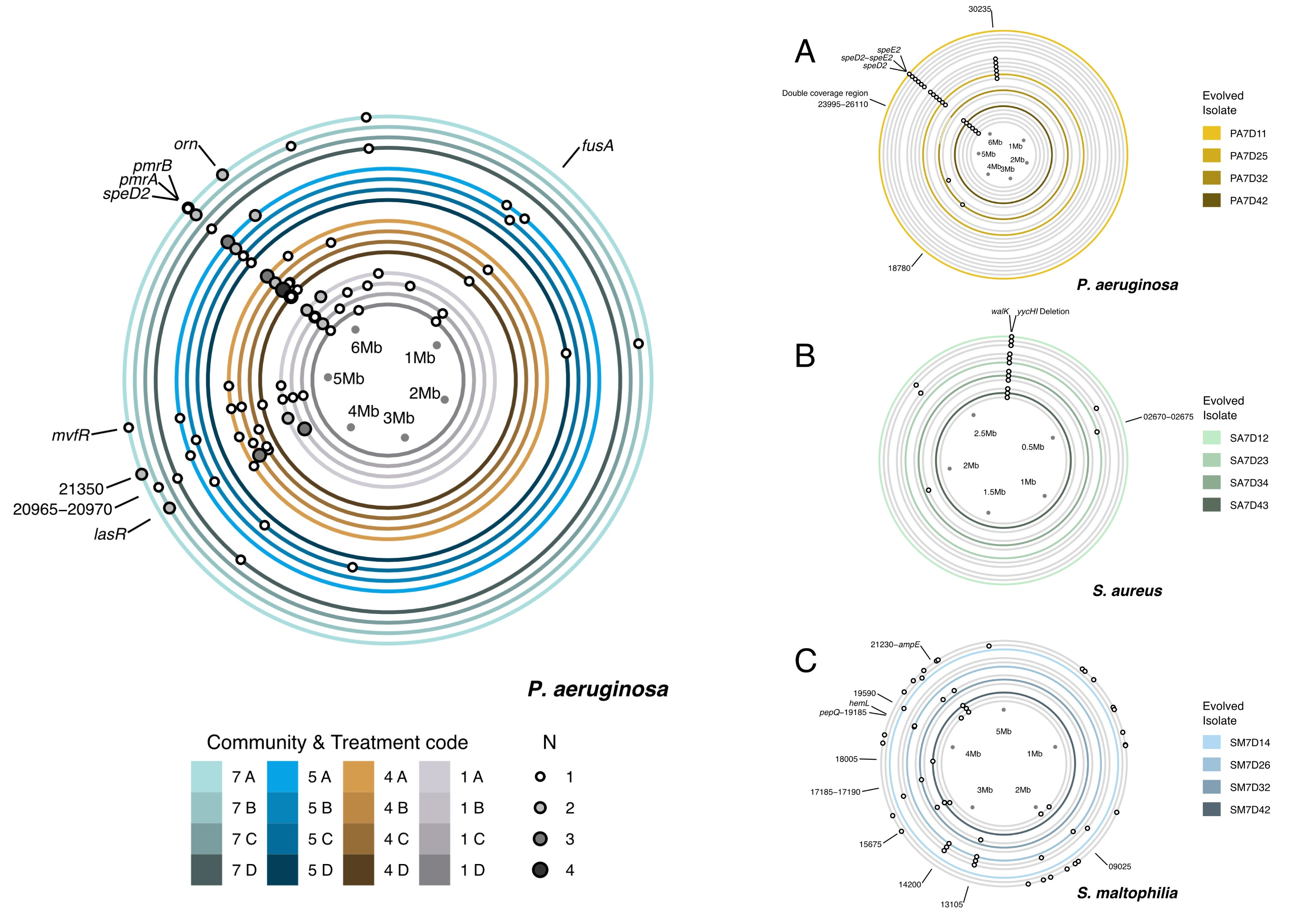
Bacteria (eg,
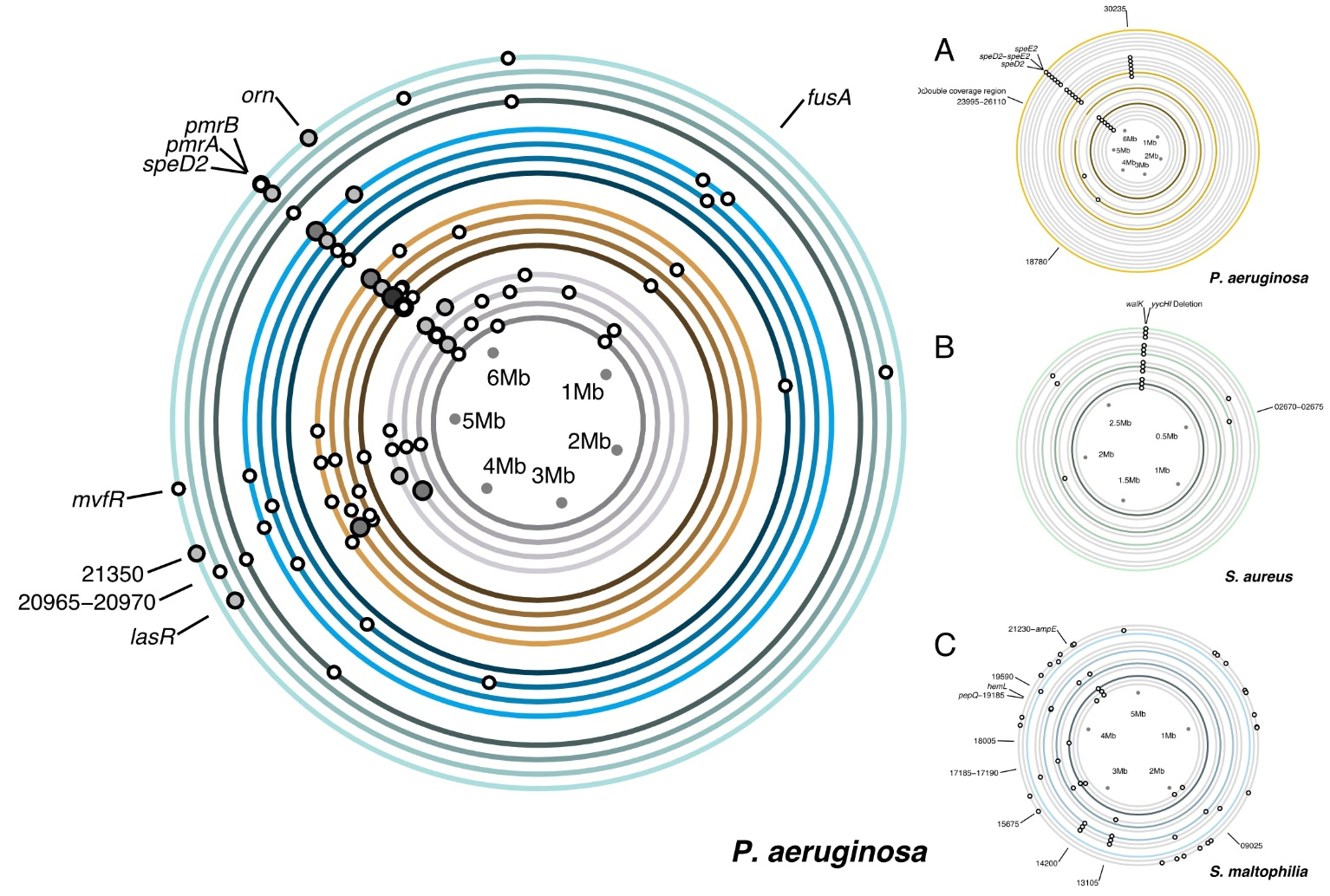
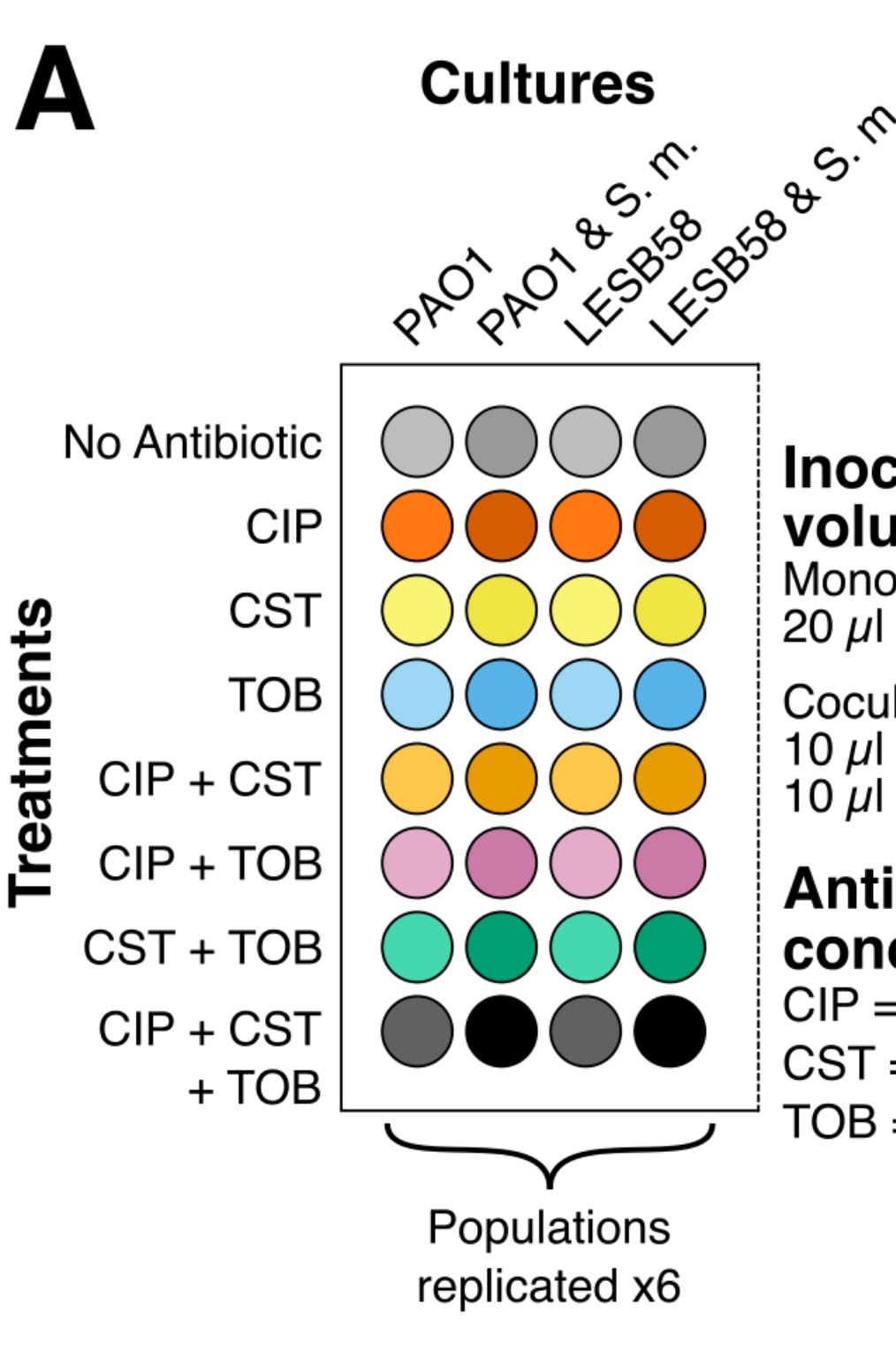
Pseudomonas aeurginosa) have circular genomes, and here I've shown the locations on the genome that developed mutations as a result of the experiment that I'd performed. The experiment had two factors each with multiple levels, community (here, 1, 4, 5, & 7) and treatment (A, B, C, & D), and showing the combinations of these two levels was the primary challenge for this set of figures.
- Distinguishing between the communities and the treatments.
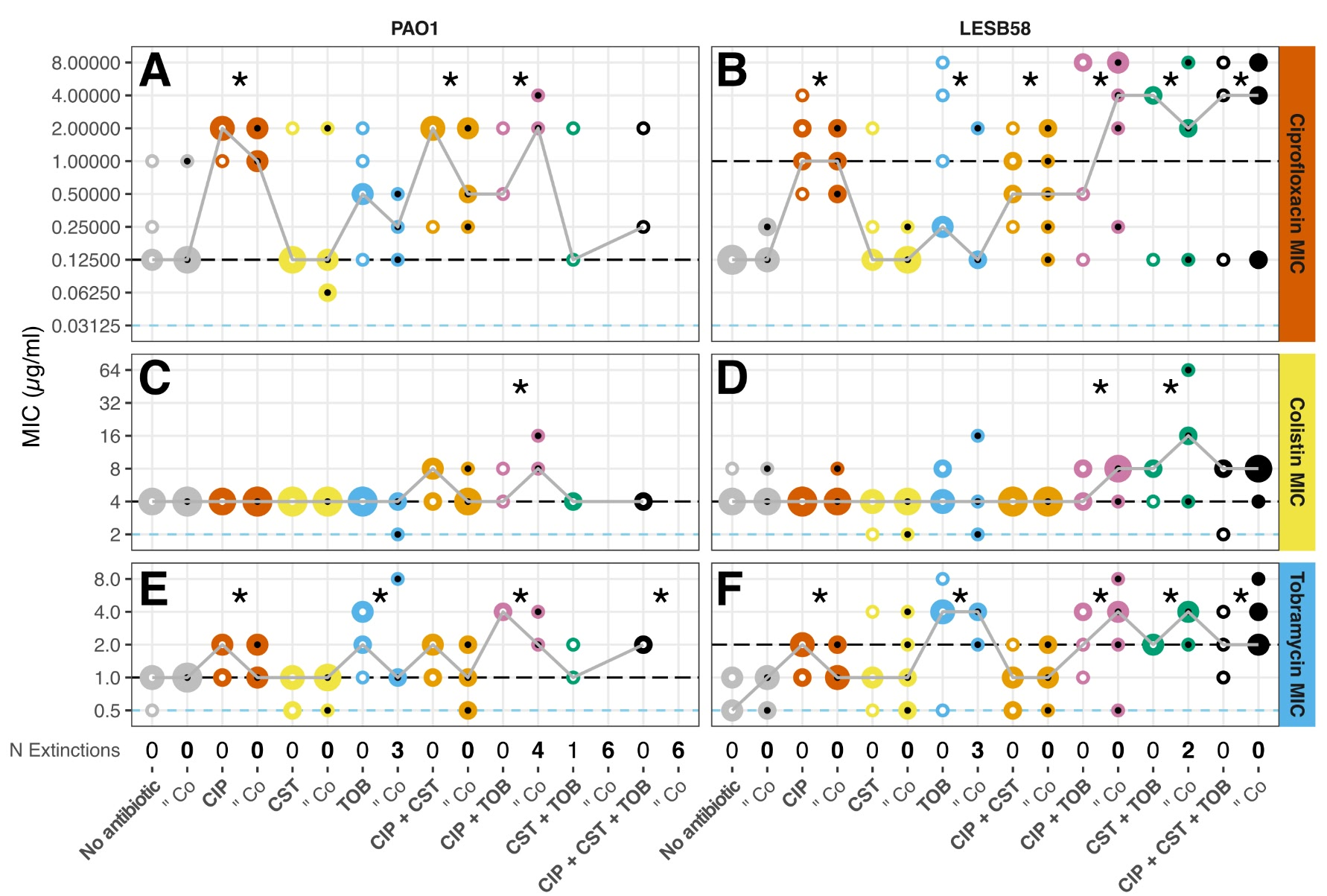
- I chose hues (with some additional spatial separation) to show communities, as this was the primary factor I was interested in, and used values of each hue to show treatments. The basic colour scheme was similar to, but distinct from, the colour scheme used to distinguish the species (in panels A, B, and C), to give a unified look.
- Logical reading order.
- I made the focus of the experiment, community 7, the largest ring as the increased circumference gives the greatest separation between the mutations. When read left-to-right, the outer ring also comes first. Similarly, within each community I chose a left-to-right arrangement of treatments.
Darkening the values towards the centre of the circle also gives a feeling of three-dimensionality, where lighter rings are higher than darker rings. This also reinforces the levels of the community factor: community 1 consisted of a single species (forming a controlled basis for my observations), communities 4 and 5 consisted of two species each (upping the complexity), and community 7 had three species (the culmination of the experiment overall).
- In panels A, B, and C, how to highlight the isolates that were used in follow up experiments.
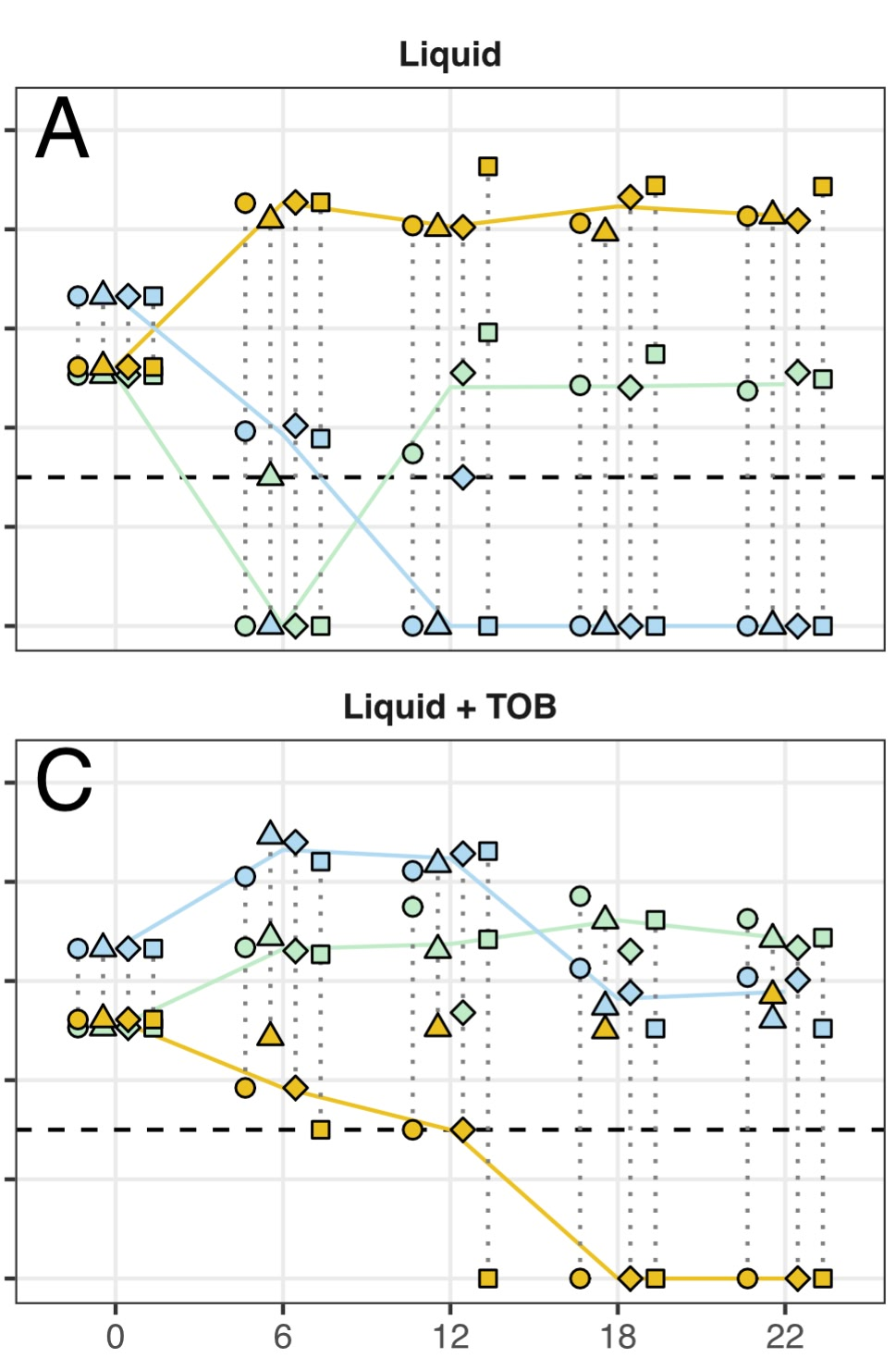
- I used the previously established colour scheme of gold, seafoam green, and sky blue for each species to highlight the isolates, and continued the value change and spatial separation to distinguish replicates.